Sixteen years ago the National Cancer Institute (NCI) and the National Human Genome Research Institute (NHGRI) joined forces to begin The Cancer Genome Atlas (TCGA), the first large-scale, global effort to characterize 33 cancer types. This data expanded our understanding of the molecular biology of cancer. Much of the focus of the original TCGA was to identify mutational profiles that can act as biomarkers to predict survival and improved treatment options. A recent study in Cell which leveraged the newly finalized survival data in the TCGA identified that mutations in tumor samples only accounted for 1% prognostic value with patient overall survival or disease progression — whereas copy number variants accounted for 30% prognostic prediction.
This is unsurprising as it is well known that copy number alterations can exacerbate genomic instability and worsen patient survival.
To that end — Mission Bio is proud to announce our launch of the Tapestri Solution for Solid Tumor Research this week.
This end-to-end solution enables researchers to design, prepare, run and analyze their samples to detect copy number and single nucleotide changes with single-cell resolution. With this solution, researchers can now unambiguously identify variant zygosity and mutational co-occurrence as well as detect rare cell populations and sub-clones in their solid tumor samples.
Design
We understand it can be challenging to select which genomic targets to investigate. To assist in this, we have worked with leading researchers in the solid tumor space, including our Center of Excellence member, Dr. Jorge Reis-Filho of Memorial Sloan Kettering Cancer Center, to develop a general breast cancer research panel. This panel contains 31 hotspot genes and 25 copy number variants and chromosome arm aneuploidies most relevant in breast cancer, to enable deeper insights into the development, progression and metastasis of breast cancer sub-types at a clonal level. Relevant targets were curated from multiple databases such as TCGA and COSMIC among others. Selecting the most frequently mutated genes down to 0.5% VAF, the targeted amplicons will detect focal amplifications, deletions or total loss of zygosity. We have also developed a similar panel for Glioblastoma Multiforme researchers. Mission Bio has utilized a similar approach in helping research groups design panels for assaying pancreatic [1], NSCLC [2], and found genomic drivers of resistance in multiple pre-clinical models of NSCLC, Pancreatic, CRC and Melanoma patients treated with KRAS G12C inhibitors [3]. Combined these results show that detection of DNA changes at a single cell level reveal previously unknown mechanisms of tumor evolution and evasion.
Prepare
We have optimized our nuclei prep protocol to ensure improved sample output and cell viability for fresh frozen solid tumor samples. This protocol has been wet lab-tested and utilized to generate the results in clinical breast cancer samples displayed below.
Run
The Tapestri Platform already supports solid tumor workflows to flow both nuclei or fully dissociated cells.
Analyze
The most significant improvement in this launch is our new copy number analysis pipeline that enables efficient analysis of copy number variants. This tool, currently deployed through Mission Bio’s expert Field Application Scientist team, can determine and depict gain and loss of copy number at a single-cell level.
As seen in the figure below, the tool was able to identify gains and losses in a clinical breast cancer sample (DCIS), at both the chromosome arm and gene level.
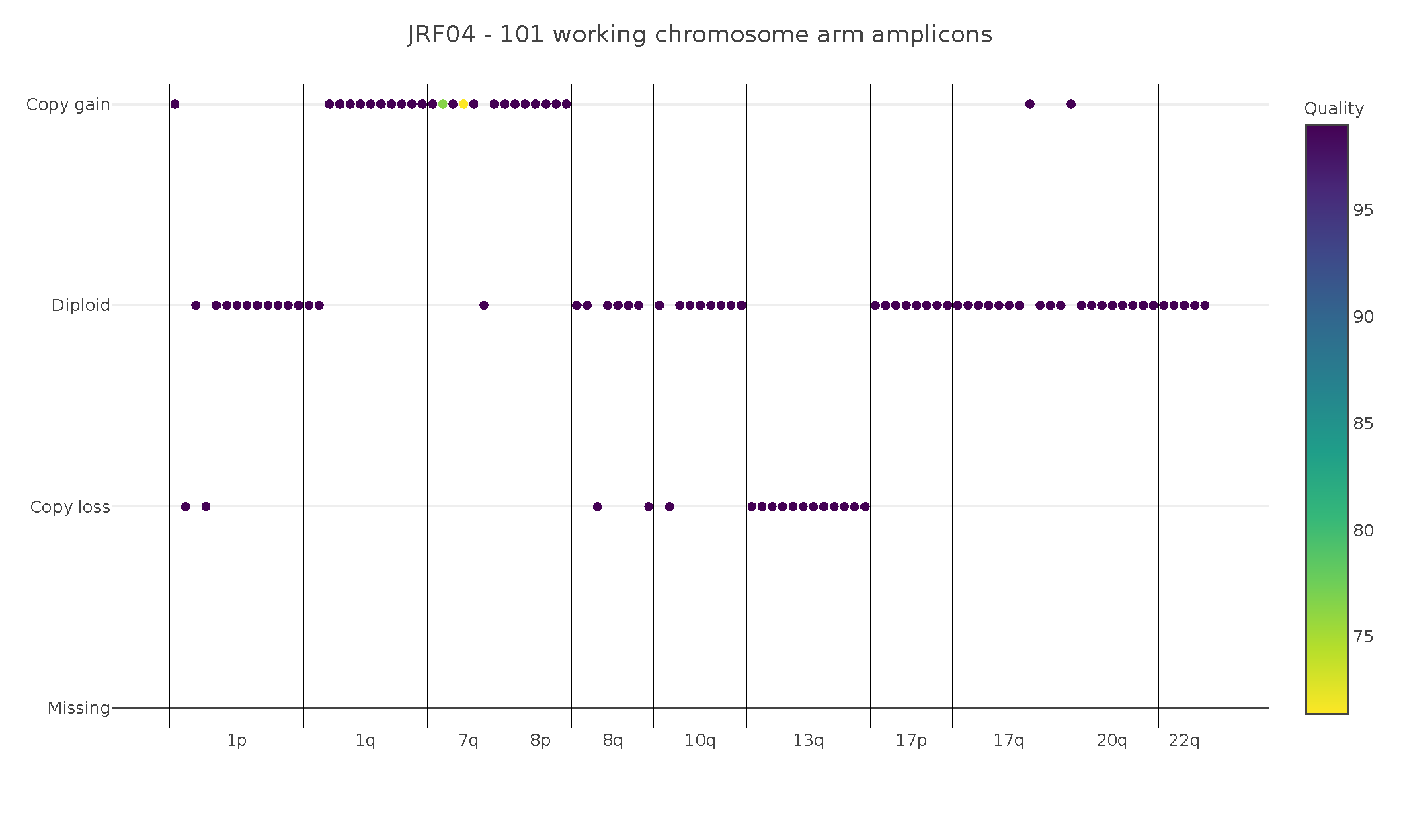
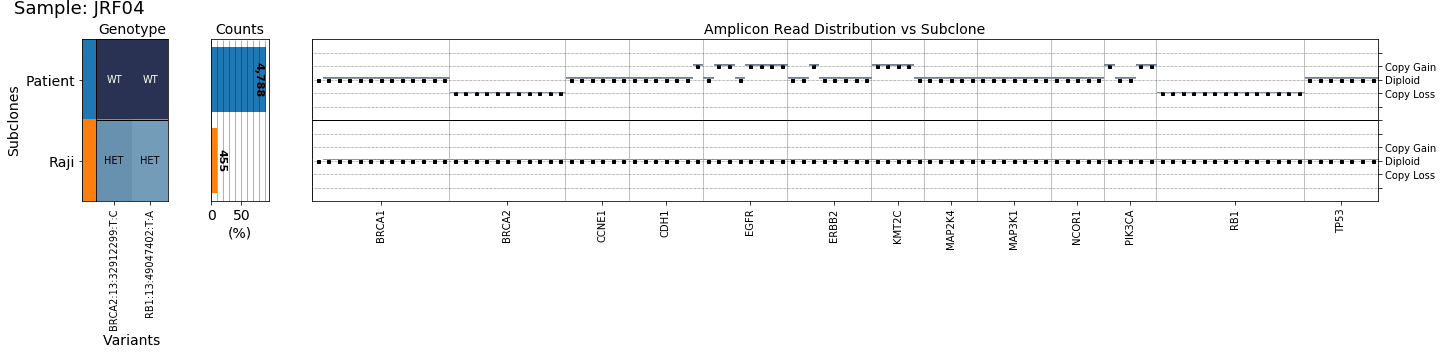
This enabled us to identify that the patient sample had a gain of the EGFR gene, commonly amplified in around a quarter of all breast cancers and loss of both BRCA2 and RB1, common tumor suppressor genes and loss of which is associated with poor prognosis.
Our mission at Mission Bio is to eradicate cancer. To do so, we must understand tumors at a clonal level. We are incredibly honored to enable cancer researchers to assess clonal architecture, uncover resistance mechanisms and delve further into the biology of disease than ever before.
For more information and to hear fascinating work leveraging the Tapestri technology, please visit us at the upcoming AACR meeting, either virtually or in-person, or get in touch with us here.



