Myeloid Published Panel
Morita, K. el al. Nature Communications (2020)
Tapestri Single-cell DNA Myeloid Published Panel was designed by the Koichi Takahashi Lab at MD Anderson Cancer Center and covers many of the recurrent gene mutations in myeloid malignancies. This panel is featured in a study published in Nature Communications, “Clonal evolution of acute myeloid leukemia revealed by high-throughput single-cell genomics.” Researchers uncovered the landscape of AML clonal architecture, revealed clonal relationships of AML driver mutations, and reconstructed the mutational history of driver genes in 154 AML samples from 123 patients. In 26 of these patients, they simultaneously examined genotype-to-phenotype correlation at single-cell resolution. In addition, scDNA-seq of longitudinal samples in 15 patients allowed illustration of clonal evolution in response to therapeutic selective pressures.
Panel Details
Tapestri Single-cell DNA Myeloid Published Panel (Takahashi, MDACC)
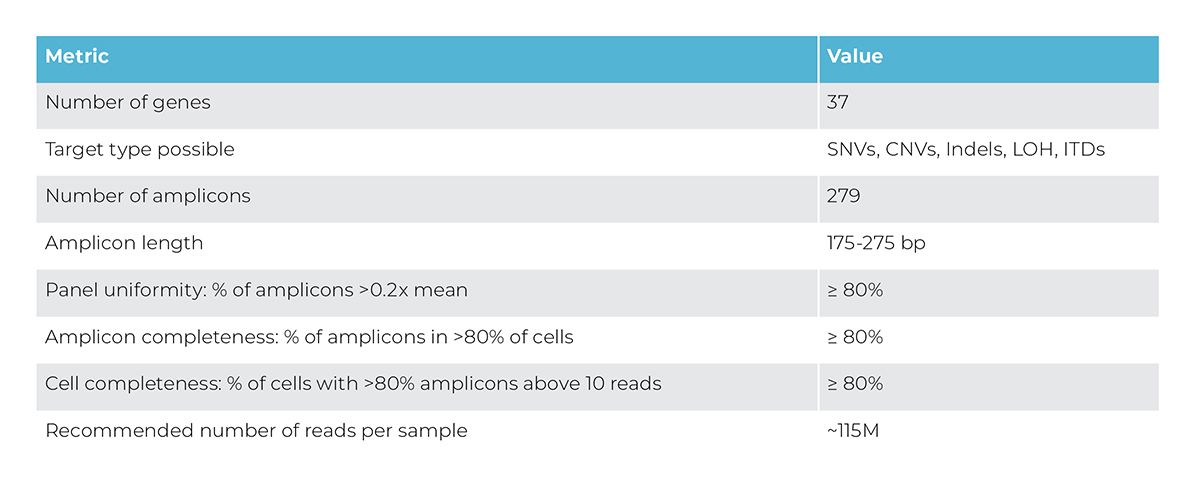
Advance your understanding of the clonal heterogeneity and evolution underpinning myeloid malignancies by targeting 37 genes with 279 amplicons for single-cell sequencing. The Myeloid Published Panel is designed to target the most commonly mutated hotspots in myeloid malignancies.
View and download panel content and coordinates using Tapestri Designer

Want to modify this panel?
Use Tapestri Designer to add or remove content from existing catalog panels and make this panel your own with content that is relevant to your research. Or start from scratch and upload your human (hg19) or mouse (mm10) targets using gene names, genomic coordinates, or SNV IDs. Target anywhere in the whole human or mouse genome and advance your research with targeted single-cell DNA sequencing.